A forked implementation of lamassemble 12, 13 ( ) was found suitable for assembling multiple long sequencing reads, 14 meeting the requirements of such sequence analysis. In the absence of a template, a consensus of all the input reads is generated to serve as a template reference to guide assembly. Prior to contig assembly, Basic Local Alignment Search Tool Plus (BLAST+) 2.90 11 is used to locally map the input reads to the template sequence. An arbitrary noise threshold ( n o i s e t h r e s h o l d) of 0.8 is used for assembly. First, read quality (described in the Supplemental Data) and trimming are performed. The ab1 traces are plotted using the D3 graphing library ( ).Īn overview of the contig assembly process is shown in Fig. YAQAAT is written in Python 3.7 using Flask ( ) and Bootstrap 4.4.1 ( ) web application frameworks with Biopython 10 and jQuery 3.4.1 ( ) libraries. 1 B) without the need for using additional web tools or software, allowing a seamless online sequence analysis experience. To allow online direct verification, translated sequences in 3 reading frames together with chromatogram peaks of mutant bases are also shown directly in the web server ( Fig. 1 A) to support quality control as well as detection of mutations. 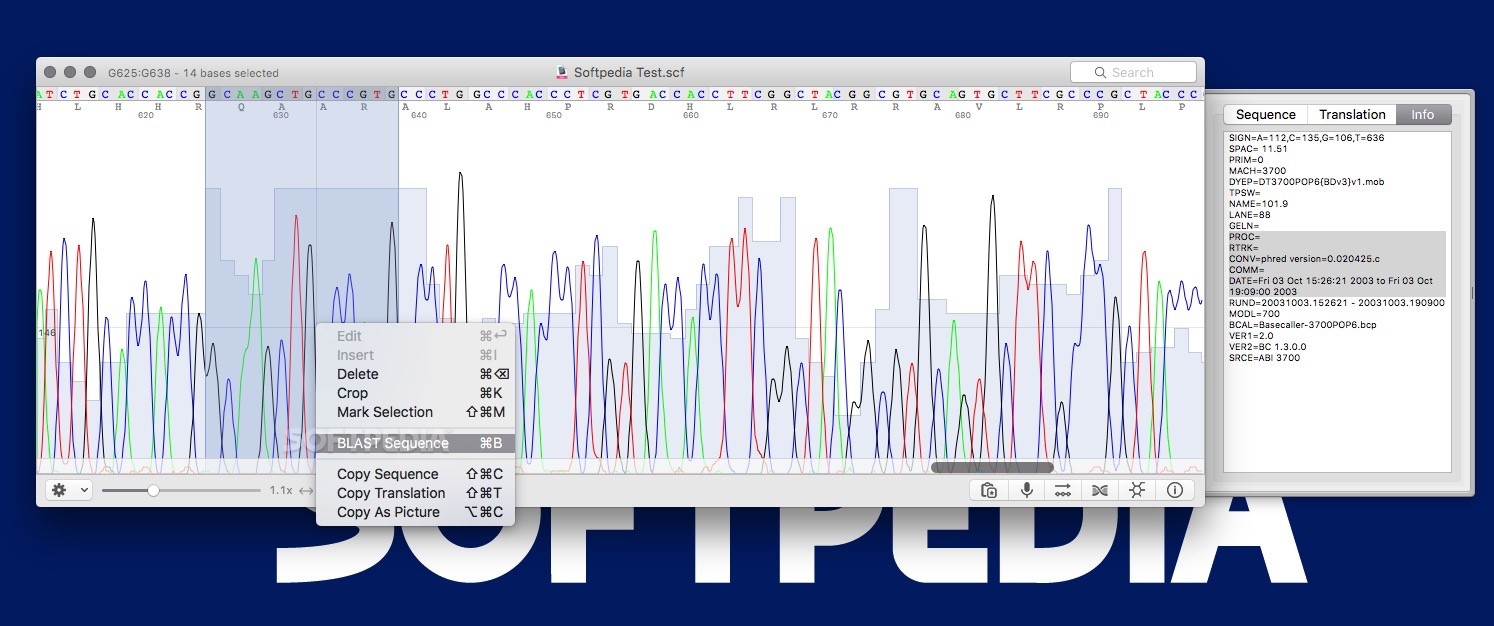
In addition, it can automatically detect single nucleotide polymorphisms/insertions or deletions when provided a reference template sequence ( Fig. To increase automation and allow analysis from any location with Internet access, Yet Another Quick Assembly, Analysis and Trimming Tool (YAQAAT) server was built to perform automated batch trimming and contig assembly not only on ab1 but also on FASTQ files. Some still require manual assembly, making the analysis of multiple sequencing samples tedious and inefficient. Many of these software programs for Sanger sequencing require stand-alone installations on the desktop/laptop, typically restricting such analysis to office spaces. To facilitate Sanger sequence analysis, many tools have been developed over the years, even as smartphone apps. 7 It is also routinely performed in general molecular methods in genetic engineering to generate. Sanger sequencing is used in primer walking, mass sequencing of error-prone PCR mutant libraries for protein engineering, 2, 3 in vitro reverse-transcriptase fidelity assays, 4 – 6 and prediction of host deaminases RNA editing sites. 
Although the capillary-based Sanger method is dated, it is still the gold standard given its superior accuracy and reliability over the second- and third-generation sequencing technologies ( i.e., next-generation sequencing, single-molecule real-time sequencing, etc.). The Sanger method, first developed by Frederick Sanger, 1 is widely used to generate accurate reads from homogenous samples economically. FASTQ sequencing reads de novo with automated trimming and scanning of the assembled sequences for single nucleotide polymorphisms and insertions or deletions without installation of software, allowing it to be accessed from anywhere with Internet access and with minimal dependency on other software and web tools. To facilitate such online accessible sequence assembly and analysis, we created Yet Another Quick Assembly, Analysis and Trimming Tool web server for the automated assembly of multiple. Through web servers with expanded automation and functionalities, even smartphones/phablets can be used to perform complex analysis previously limited to desktops, especially if they can upload files from cloud storage. With increased data output worldwide, there is also a need for automated quality checks and trimming prior to large assemblies, along with automated detection of mutations. Even with the ubiquity of Sanger sequencing, automated assembly software are predominantly stand-alone software packages for desktop/laptop use with very few online equivalents, thus geospatially constraining sequence analysis and assembly.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
